September 7, 2021 (Huntsville, Alabama)- Members of the Felidae family, more commonly referred to as cats, come in a variety of sizes and a range of varying fur colors and patterns. Tigers with black stripes on an orange or white background roam the grasslands of Southeast Asia and India. Spotted cheetahs chase antelope through the shrubland in Africa. Jaguars with their rosette patterned coats stalk down prey throughout South and Latin America. And domesticated cats of many coat colors and patterns bask in the sunlight of living rooms across the world.
Scientists agree that hair follicle cells are the source of the black, brown, yellow, and red pigments that color hair or fur. However, when and where the process of establishing the color pattern takes place has been less clear until recently. A team of scientists at the HudsonAlpha Institute for Biotechnology identified molecules that regulate the establishment of coat color patterns in cats. The results are published in Nature Communications.
Development of color pattern
Prior studies have shown that in tabby cats stripes are evident in the fetal cat as soon as pigment is produced by cells called melanocytes within the hair follicle. After birth, newborn cats have the same coat pattern and color that they will have for their entire life. For example, cheetahs are born with the same number of spots as they will have in adulthood, the spots just grow in size. As hair falls out, new hair of the same color replaces it.
The origin of coat color is well known, but the mechanisms that initiate coat color patterns are less understood. Complex arrangements of alternating patches of light and dark hair as seen in a cheetah, jaguar, or ocelot, are called periodic color patterns. Such patterns are difficult to study because they have no real counterpart in model organisms.
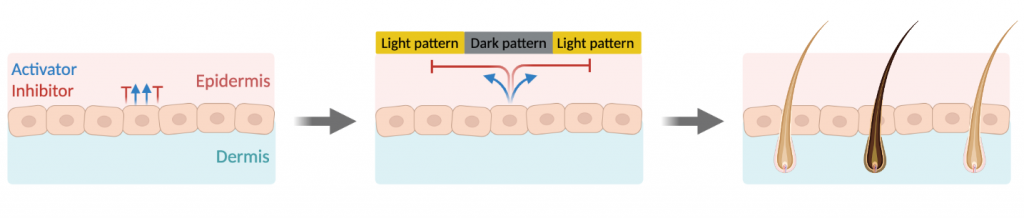
In the early 1950s, the computing pioneer Alan Turing theorized that molecules that inhibit and activate each other could create periodic patterns in nature if they travel through tissue at different rates. This theory, called the Turing reaction-diffusion mechanism, means that coat color patterns that form during development could be controlled by activator and inhibitor molecules. The activator molecules simultaneously color a nearby cell and trigger the production of inhibitors that travel to distant cells and shut off pigment production.
“The genes that control simple color variation, like albinism or melanism, are the same in all mammals for the most part ,” says animal morphology expert Greg Barsh, MD, PhD, HudsonAlpha Faculty Investigator, Faculty Chair, and Smith Family Chair in Genomics. “However, the biology underlying mammalian color pattern has long been a mystery, one in which we have now gained new insight using domestic cats.”
HudsonAlpha scientists discover gene that controls periodic color patterns
Barsh and HudsonAlpha senior scientists Chris Kaelin, PhD, and Kelly McGowan, MD, PhD, previously showed that a gene called Endothelin 3 is expressed at the base of hair follicles in tabby cat markings and plays a key role in the development of tabby pattern. Because the tabby markings are apparent in developing hair follicles, the group hypothesized that establishment of color pattern must occur at or before hair follicle development. They set out to determine exactly when, where and how patterns are established in developing fetal cats.
McGowan and Kaelin developed partnerships with several feral cat trap-neuter-release programs, collecting fetal tissue that would have otherwise been discarded during spay procedures. McGowan studied the fetal cat skin tissue and found stripe-like alterations in epidermal (skin) thickness early in fetal development. The patterns of epidermal thickness resembled tabby fur patterns in adult animals.
“Our findings from the morphological studies suggest that even before melanocytes enter the epidermis, the cells are predestined to signal for a specific fur color,” says McGowan. “By understanding the developmental window and cell type in which color pattern establishment occurs, we were able to dive deeper and discover the molecules involved in pattern development.”
Building on this new finding, Kaelin used single-cell gene expression analysis on fetal cat skin cells just prior to the time at which the thick and thin patches become apparent. Through this analysis, the team determined that epidermal expression of a gene called Dickkopf 4 (Dkk4) marks areas of fetal skin that give rise to hair follicles that later produce dark pigment. Dkk4 is an inhibitor of Wnt signaling, which helps determine cell fates and spurs cell growth during development in many animals.
“Our analysis identifies a network of molecules involved in pattern formation,” says Kaelin. “Several of the molecules, including Dkk4, are known to function coordinately as activators and inhibitors, exactly as Alan Turing predicted 70 years ago.”
Further experiments also showed that Dkk4 is linked to other color patterns in cats. In the Abyssinian cat, the apparent absence of dark tabby markings, a trait referred to as “Ticked”, accentuates the alternating color bands present on individual hairs. The team discovered that all Ticked cats carry loss-of-function mutations in Dkk4.
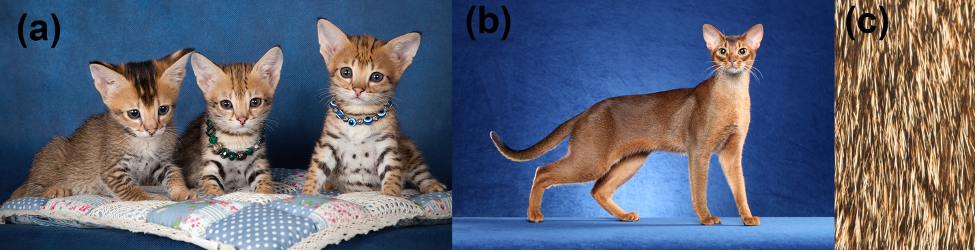
(a) A Dkk4 mutation in the kitten without the necklace causes dark tabby markings to become smaller and more numerous. (b) An Abyssinian cat with a Dkk4 mutation that is associated with an apparent absence of tabby markings. (c) Close-up view of the Abyssinian coat suggests that microscopic dark markings may be present (Photo credit for c, Martin Bahmann CC BY-SA 3.0)
“In Abyssinian cats with the Ticked phenotype, the consensus has been that there is an absence of the dark tabby markings,” says Barsh. “Based on our new findings, we propose that instead the typical tabby markings have increased in number and decreased in size to the extent that they are just not readily apparent.”
Taken together, the results presented in this study confirm a direct role for Dkk4 in cat color pattern establishment. Coat color and pattern variation is an important platform for studying gene action and interaction. Previous work in Barsh’s lab has shown that coat-color genes are involved in many other important biological pathways, some having relevance to human health.
To learn about the history of this project, please read Barsh’s “Behind the Paper” blog post here.
Byline: Sarah Sharman, PhD, Science writer
About HudsonAlpha: HudsonAlpha Institute for Biotechnology is a nonprofit institute dedicated to developing and applying scientific advances to health, agriculture, learning, and commercialization. Opened in 2008, HudsonAlpha’s vision is to leverage the synergy between discovery, education, medicine, and economic development in genomic sciences to improve the human condition around the globe. The HudsonAlpha biotechnology campus consists of 152 acres nestled within Cummings Research Park, the nation’s second largest research park. The state-of-the-art facilities co-locate nonprofit scientific researchers with entrepreneurs and educators. HudsonAlpha has become a national and international leader in genetics and genomics research and biotech education and fosters more than 45 diverse biotech companies on campus. To learn more about HudsonAlpha, visit hudsonalpha.org.
Media Contact:
Lara Burhenn
lburhenn@hudsonalpha.org
(256) 937-8210