Plant geneticists work on some of the most pressing questions for humanity. With a changing climate and a growing population, scientists and farmers have to work together to find versatile, resilient crops to maintain a stable food supply.
People often focus on GMO’s, which we discussed in an earlier Shareable Science, but as we also pointed out in that article, genetics help with many traditional plant breeding techniques.
For example, for thousands of years, farmers have intentionally crossed their most successful plants and used those seeds for the next generation of crops.The process is called selective breeding, and it can be helped along significantly by genetics. Many important crops, from food sources to biofuels, take years to mature. That means the selective breeding process can take decades to solidify new, desirable traits like disease resistance or better water efficiency.
However, if you know which parts of the genome affect those traits, you can breed new plants then sequence them when they are young to find out which ones carry the combination of traits you need. Selective breeding with the assistance of genomics lets the process move many times faster.
In order to facilitate that rapid improvement, you need a point of comparison, which is why agriscientists develop what are called reference genomes. The name sums them up neatly. They are genomes you can reference to see where a specific variety of plant has deviated from the standard, letting you know whether it has acquired desirable or undesirable traits in the breeding process well before the plant reaches maturity.
The HudsonAlpha Institute for Biotechnology’s Genome Sequencing Center (GSC) is a global leader in generating these types of reference sequence. They specialize in applying genomic techniques to understand how plants function in response to environmental factors like water, temperature, fertilizer and harmful pests. Led by faculty researchers Jeremy Schmutz and Jane Grimwood, PhD, their team has essentially been part of the development of half of all plant reference genomes known to date.
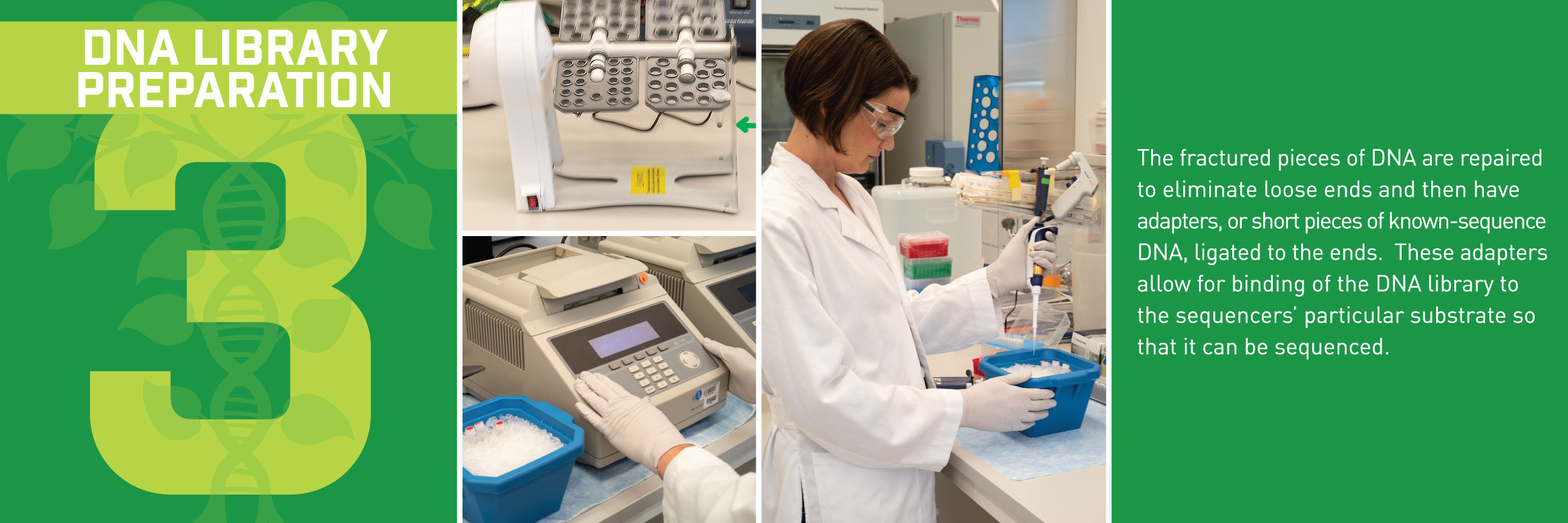
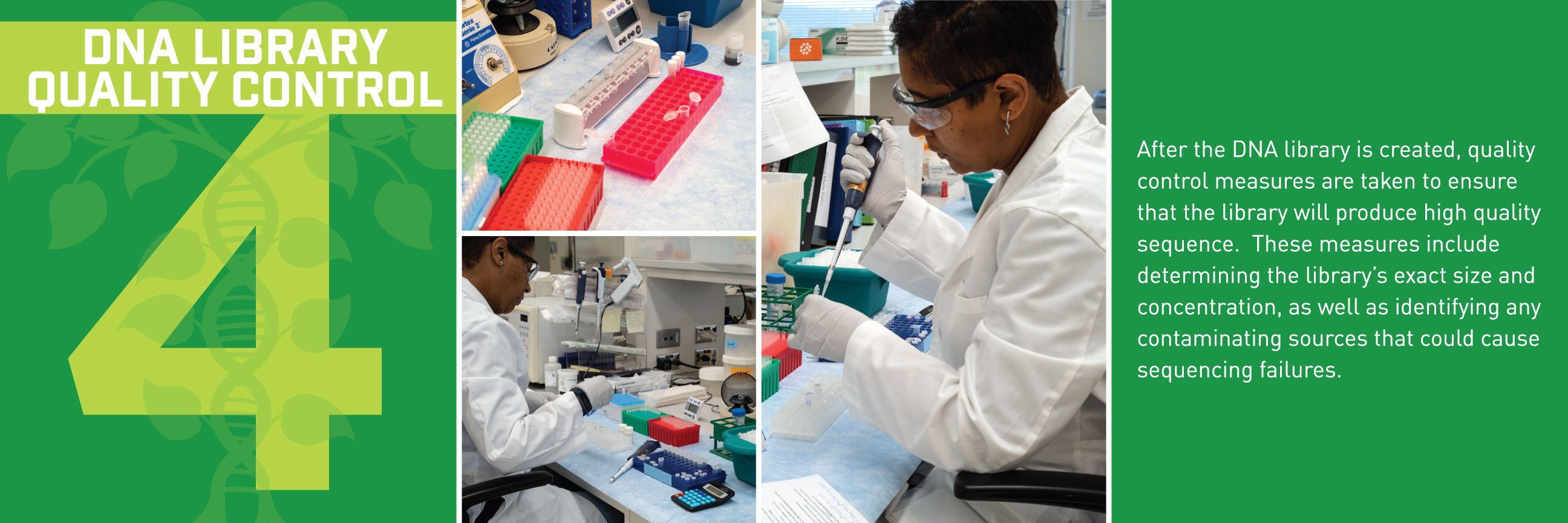
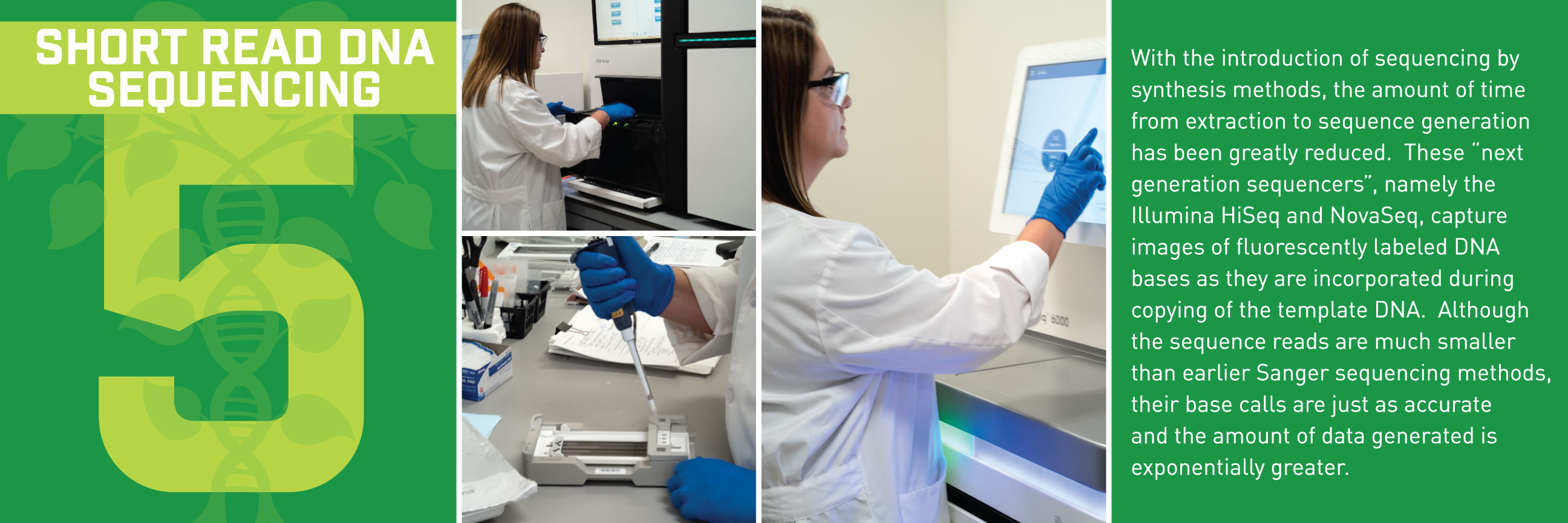
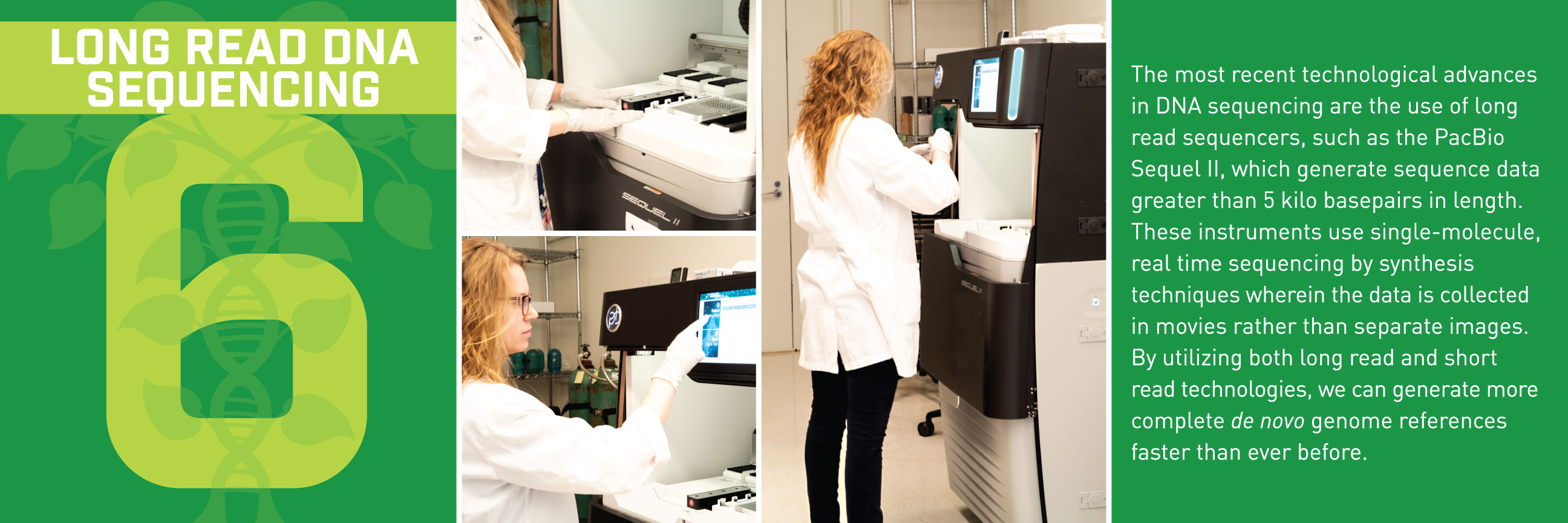
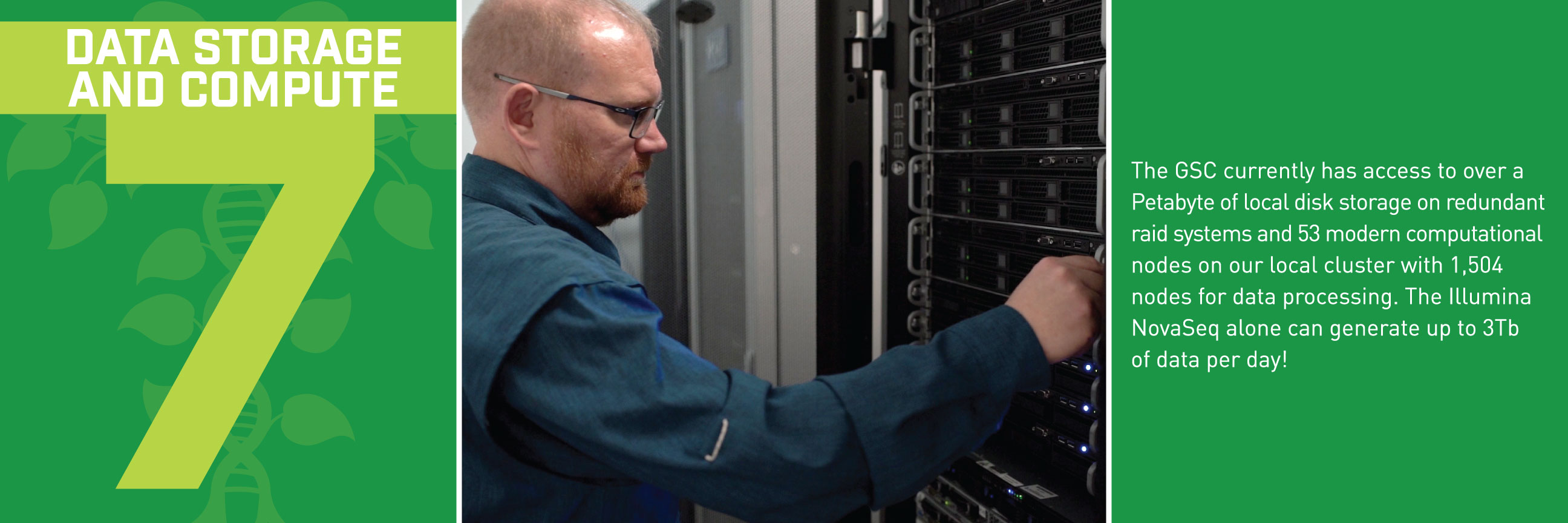
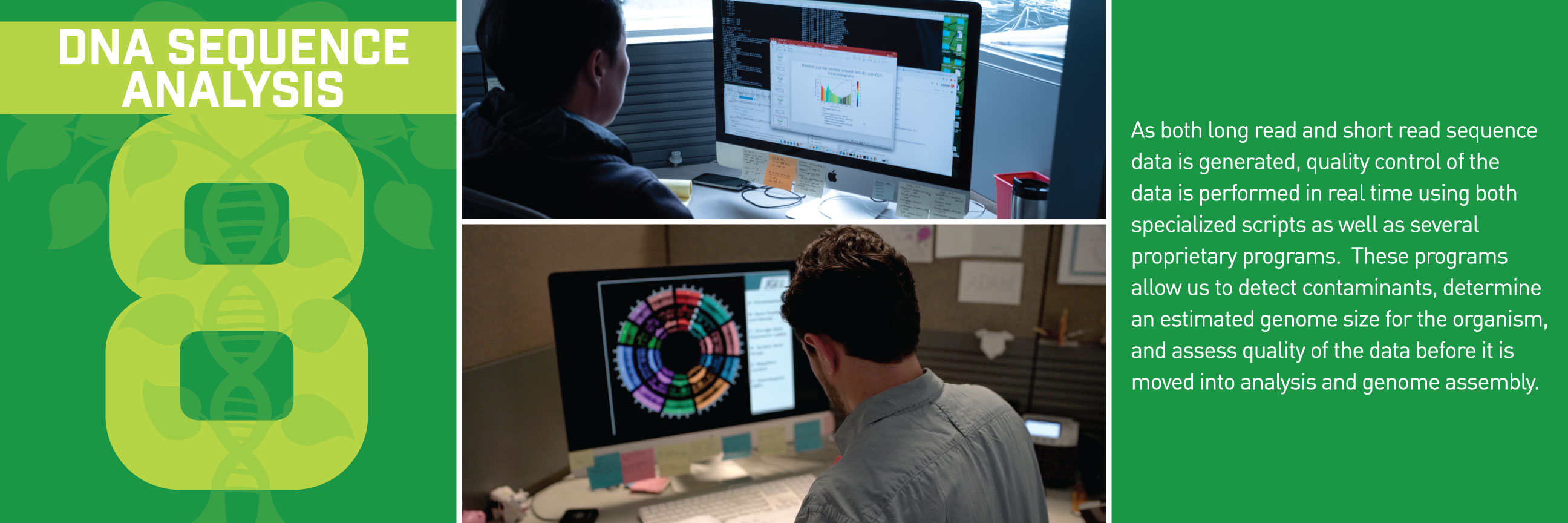
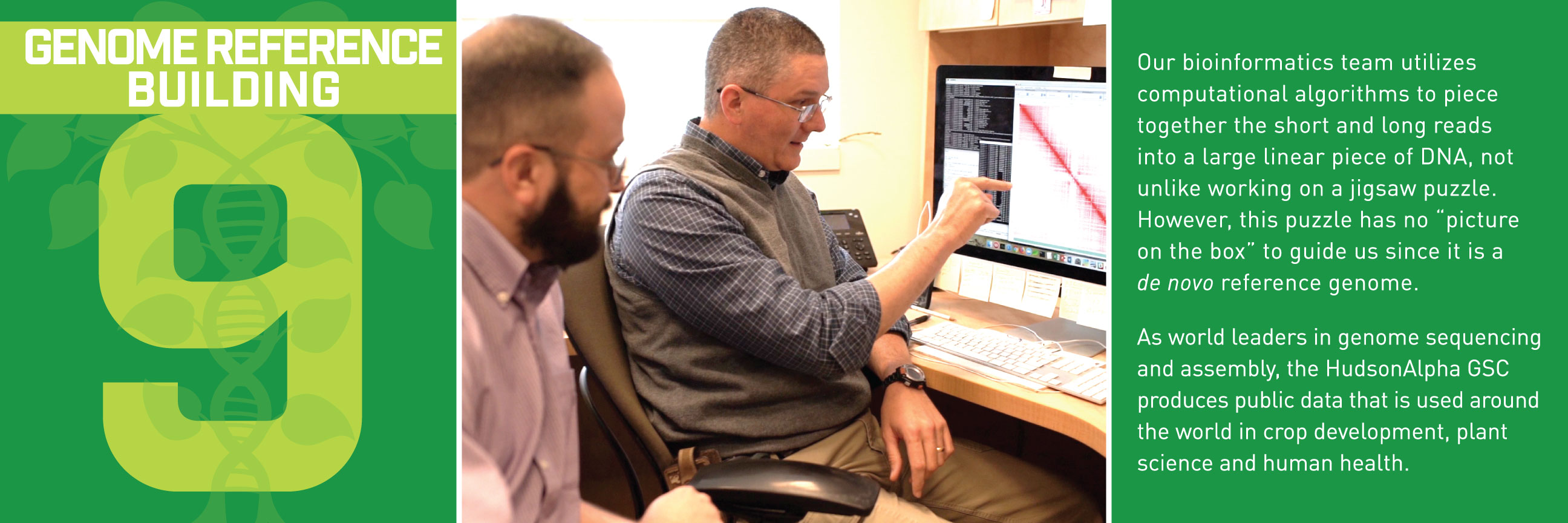
Typically, we’d talk about some new application of these reference genomes, but this week, we wanted to break down how the sequences get built in the first place. Thanks to the communications department at HudsonAlpha, we have a beautiful (and informative) step-by-step guide for making reference genomes.
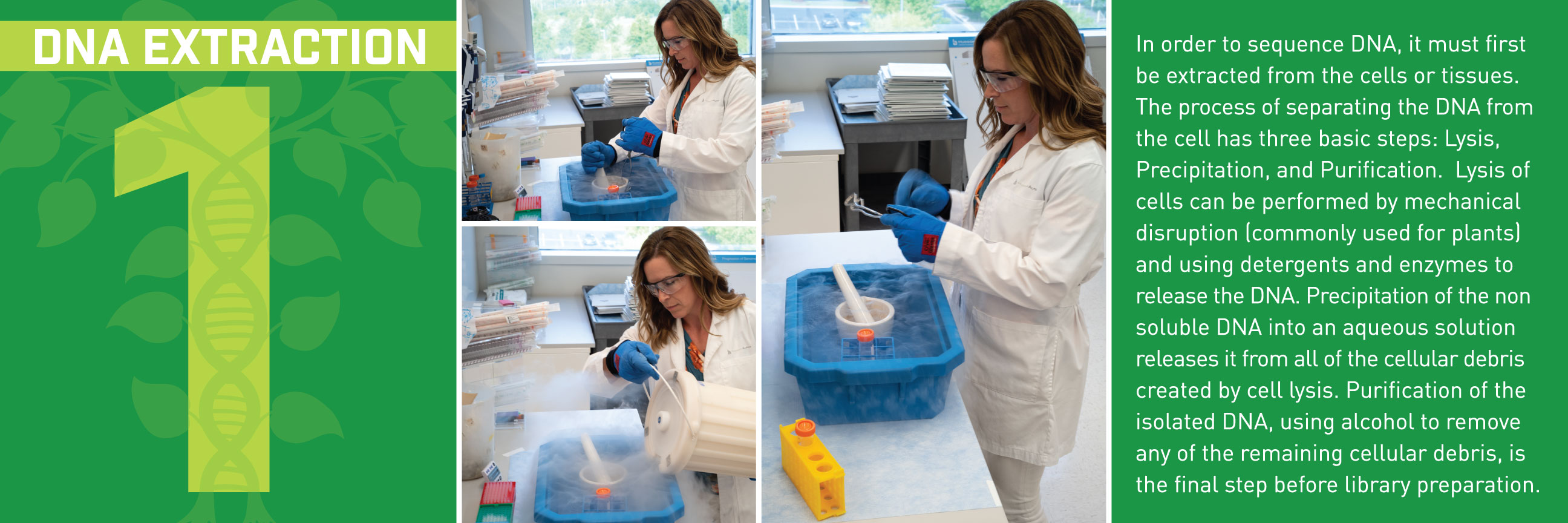
You start by pulling DNA out of plant cells.
HudsonAlpha’s GSC has generated a number of important reference genomes for everything from the American chestnut to cotton. The Institute has created reference sequences for popular crops like cacao, which is critical for maintaining the world’s chocolate supply. But more importantly, the cacao reference genome will help to ensure cacao can be produced in the parts of the world that currently rely on the crop for income. The team has also created reference genomes for less broadly known crops like switchgrass, which help us understand plant mechanisms for drought resistance.
Enhancing the world’s crops serves an important and noble purpose. It helps secure food supplies and develop potential energy sources, while also creating avenues to make the crops we grow more productive and resilient.
To schedule a media interview with Dr. Neil Lamb or to invite him to speak at an event or conference, please contact Margetta Thomas by email at mthomas@hudsonalpha.org or by phone: Office (256) 327-0425 | Cell (256) 937-8210
Get the Latest Sharable Science Delivered Straight to Your Inbox!
[gravityform id=19 title=false description=false ajax=true][wprpw_display_layout id=8]